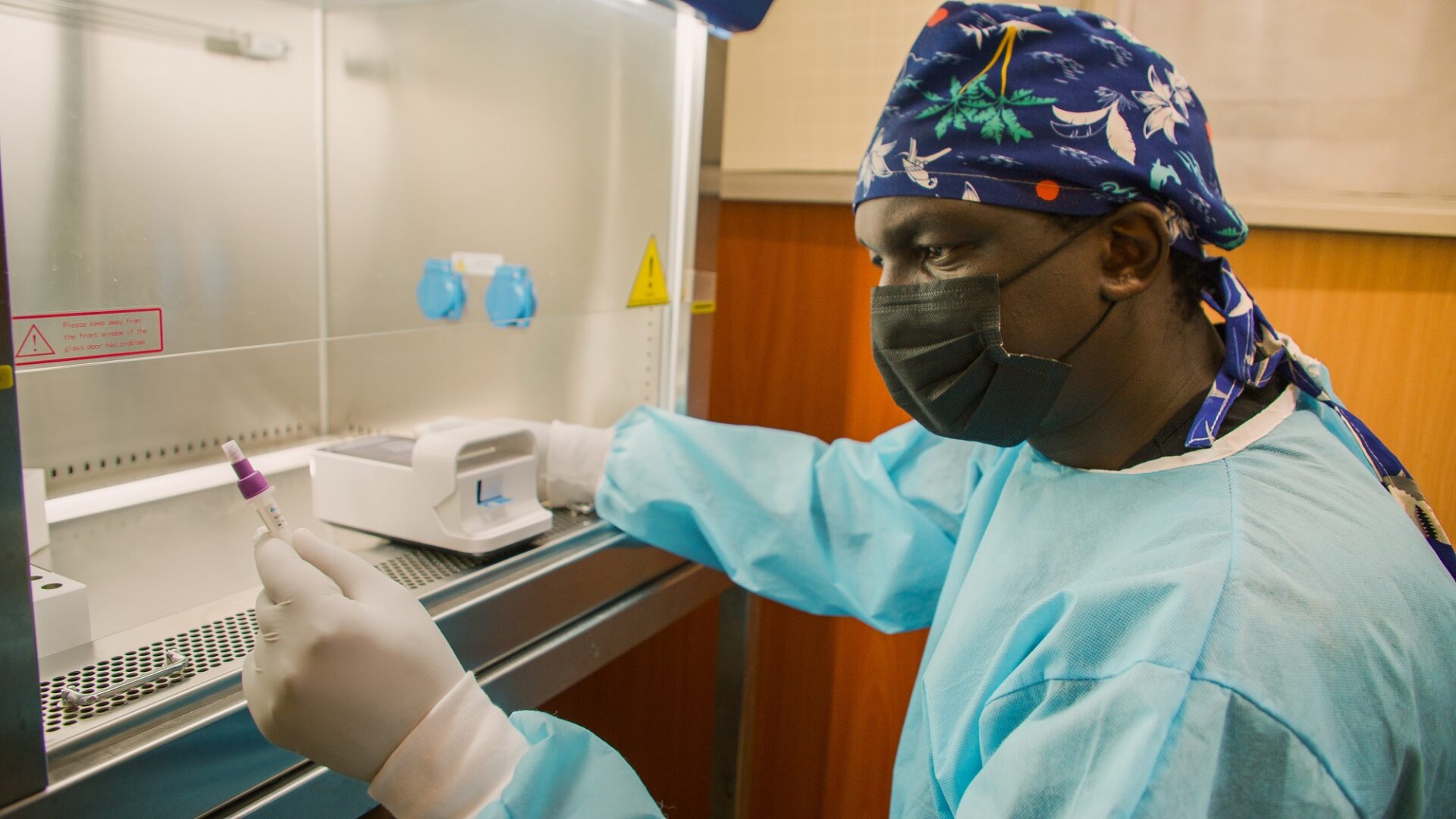
Happy 2022! We turned the page of the calendar, but for many people around the world, life feels eerily similar to the early days of the pandemic as schools closed, restaurants shifted to curbside pick-up and take-out only options, and testing kits were only available to those who staked out online retailers in the wee hours of the morning.
But we know that this is not 2020. We have effective vaccines, testing has ramped up significantly, and healthcare workers in many parts of the world have much more experience with treating this disease. And, the approval and preliminary roll-out of new treatments provides additional hope on the horizon.
We’ve all increased our knowledge of public health terminology over the past two years. But 2021—and now 2022—have taught us about variants. We heard about the alpha variant that was first detected in Britain, but it wasn’t until the delta variant that we all perhaps tuned in a bit more closely to how viruses change over time.
We want to take you behind-the-scenes to the world of scientists and researchers who study the genomic sequences not only of viruses like SARS-COV-2, but also bacteria and other microorganisms that can cause disease. And, we want to highlight the work of the Africa Pathogen Genomics Initiative, which is ramping up capacity to conduct genomic surveillance across the African continent.
We recently spoke with Dr. David Blazes, a Deputy Director on the Gates Foundation’s Global Health team who leads the foundation’s work on Genomics, Epidemiology, and Modeling, and Dr. Jennifer Gardy, a Deputy Director whose scientific background is in genomic surveillance and who now oversees how this technique is rolled out as part of the fight against malaria.

Gates Philanthropy Partners: Genomic surveillance and genomic sequencing are not something most people in the world have heard of until this pandemic. Can you help us understand why this field is important beyond what we know about COVID-19 variants like delta and omicron?
Dr. Blazes: Let’s start out by defining some terms. When we’re talking about pathogen genomic surveillance, this is a field of research that helps those of us who work in public health to detect and characterize emerging diseases in a population, like how SARS- corona virus 2 (SARS-CoV2) was detected as a new pathogen—as well as endemic diseases, like polio in Pakistan and Afghanistan, where we’re able to see where the virus is still circulating. When we’re using the term genomic sequencing, this is the use of various scientific technologies and methods to look specifically at the DNA and RNA of pathogens. To use this in COVID-19, there are researchers that are studying lab samples of SARS-CoV-2, taken from patients, to determine changes in the DNA and RNA, which is how new variants are identified and characterized. This is the sequencing. The gathering of those samples to test and then the study of where those variants are appearing in the population is the surveillance.
Dr. Gardy: COVID-19 has certainly drawn attention to this field, but this work has been around for a while and has become increasingly common over time. About a decade ago it was mostly one-off studies of individual disease outbreaks, then the Ebola epidemic from 2014-2016 was one of the first major disease events that genomic surveillance helped us to track and understand. And it’s not just viruses: everything from the DNA of parasites that cause malaria and neglected tropical diseases to the bacteria that cause infections like whooping cough can be sequenced.
No matter the pathogen, what’s most important is what we as researchers and public health experts learn from that data—how we turn that data into action. In our work on malaria, for example, we can use sequencing and surveillance can tell us so many things. They can tell us where malaria parasites and where mosquito vectors are and how their populations are changing over time. We can also look at where we’re seeing drug-resistant parasites or insecticide-resistant mosquitos so that we can adapt treatment and control measures to try to stay one step ahead of infections.

Gates Philanthropy Partners: We heard a lot about South Africa detecting the omicron variant—and it seems that there are only certain countries that are quite well-equipped to do this type of research. Is that true?
Dr. Blazes: Yes and no, and this is rapidly evolving! There are well-established—and well-funded—labs in places like the UK and some other sites in Europe, the US, Singapore, Australia and South Africa that created this field. One of the initiatives we’re funding and quite excited about is the Africa Pathogen Genomics Initiative, which launched in 2020 as a joint initiative of the Africa CDC and member states of the African Union. This initiative aims to ramp up some of the already excellent work being done on the continent and build capacity in countries where there are gaps in genomic sequencing and surveillance.
When we think about the expanse of the African continent and the diversity of populations—and, unfortunately, the impact of infectious diseases like malaria, cholera, tuberculosis, and HIV, we see this initiative as game-changing. With high-quality genomic epidemiology that is able to inform public health experts and other decision-makers about where diseases are circulating, we increase the knowledge and tools available to monitor cases, control disease spread, identify effective interventions and determine the optimal prevention and treatment measures.
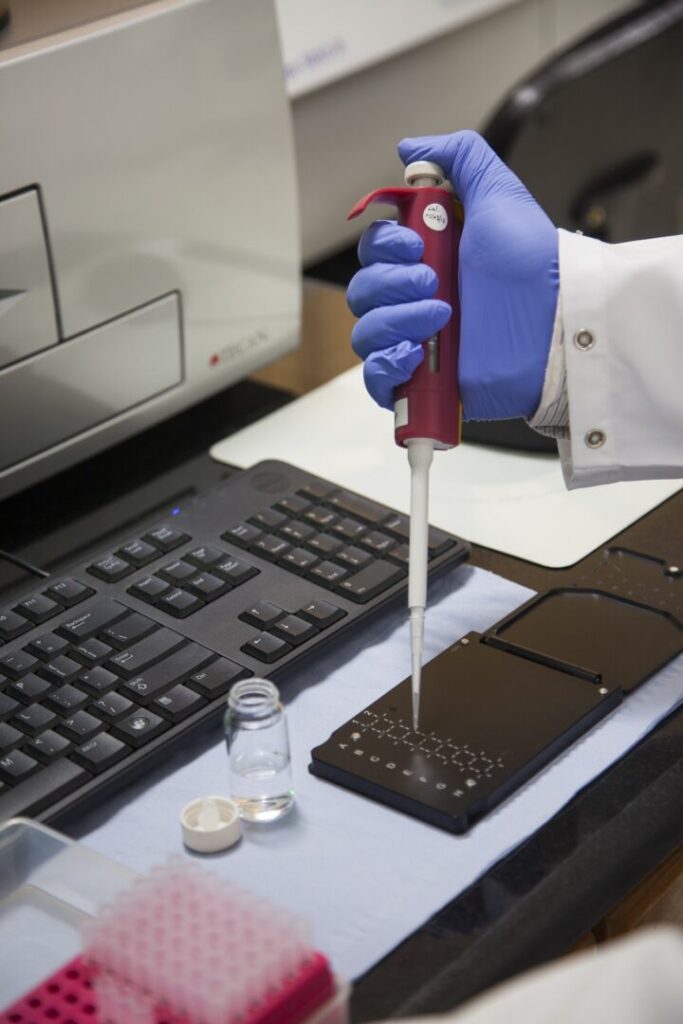
Dr. Gardy: One thing to add here is that many countries on the continent—and elsewhere in the world—face resource constraints when it comes to public health. Traditional surveillance can be very expensive, time-consuming, and requires a lot of specialized personnel. But by analyzing DNA from, say, a used rapid test for malaria or a swab from a COVID test, we can get a lot of useful information affordably, quickly, and with a single analysis. Genomic sequencing won’t ever replace traditional surveillance, but it’s a great complement that can fill some critical gaps.
Our goal in funding this initiative (and the other groups we fund to work on individual genomic surveillance projects around the world) is to democratize the access to and use of these technologies globally so that everyone can benefit from this sort of enhanced surveillance. As David noted, this approach is routine in many parts of the world, but when we’re working with infectious diseases, a disease anywhere can easily be a disease everywhere, so we see the need for these technologies to be easily used in as many countries as possible.
Gates Philanthropy Partners: The Africa Pathogen Genomic Initiative is quite new. Is this a result of the pandemic?
Dr. Gardy: Having worked in the genomic surveillance domain for a decade before I joined the Gates Foundation, I can definitely tell you that pre-pandemic, it wasn’t always easy getting funding for this sort of work. But the pandemic has certainly accelerated public health agencies’ interest in this approach and forced a rapid ramp-up in capacity around the world. This is true across Africa too. Pre-pandemic, there was some great critical mass around genomic surveillance and some wonderful collaborations around building regional networks and increasing training efforts, and the pandemic really kicked those into high gear.
Dr. Blazes: We’ve really seen this initiative take off in the past year. For example, at the start of 2021 there were about 5,600 sequences of SARS-COV-2 in existence and by the end of 2021, there were over 60,000. The initial plan was to phase-in labs in 20 countries over four years, but the initiative has already expanded to 45 countries. Some of those aren’t fully up-and-running and are still sending samples to labs with more capacity in Nigeria and South Africa, but that’s incredible growth in just a year. COVID-19 really accelerated that growth, which hopefully can be leveraged for many other diseases.
Gates Philanthropy Partners: What’s on the horizon for this work—in Africa and elsewhere?
Dr. Gardy: For our malaria genomic surveillance work, we’ve invested in “anchor” labs in Senegal, Tanzania, Mozambique, and Uganda that not only have the capacity to do big, national surveys of malaria parasites, but that can also act as partners and trainers for other countries who are just starting to set up malaria genomics projects. We’re eager to keep growing this South-South training model to encourage knowledge sharing across the continent with learnings and expertise specific to those geographic contexts.
Dr. Blazes: In the short-term, these labs will continue to be plenty busy with COVID-19 response, particularly as vaccine roll-out in many countries is disappointingly slow and creates the opportunity for more variants to emerge. In the medium-term, we hope that the initiative is able to also address cholera, yellow fever, tuberculosis, and other diseases. Longer term plans include building links between genomic surveillance and local manufacturing of diagnostics, medicines and vaccines. We’ll also be eager to see the types of networks that are built between labs—as well as ministries of health and the Africa CDC.
Thank you, Dr. Blazes and Dr. Gardy for sharing your expertise in this area. When we hear about variants related to the current pandemic we’ll now know a bit more about the science behind those discoveries as well as how those labs are contributing to the fight against many other pathogens.

